
M.S. - 1993 - Seoul National University
Ph.D. - 2000 - University of Wisconsin-Madison
Postdoc. 2001-2005 - University of California, Berkeley
Not Accepting Students
Proteins are dynamic molecules. Even under native conditions, they do not adopt a single static conformation. Rather, they access many different conformations in their native state ensemble. This native state ensemble includes small fluctuations around the native conformation, partially unfolded forms, and even globally unfolded forms. The distribution of these conformations and the kinetic barriers between the conformational states define the conformational energy landscapes of proteins. My research interest is investigating conformational energy landscapes of proteins and deciphering the relationship between the energetics of proteins and their biochemical functions, such as catalysis, signal transduction, and ligand binding. We use proteolysis as a major tool to probe protein structures and dynamics as well as conventional spectroscopic methods. We also use proteomics extensively for investigating energy landscapes of proteins on a system level.
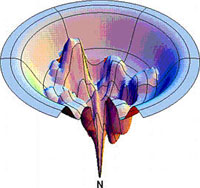
Conceptual representation of a conformational energy landscape of a protein (Dill & Chan, Nature Struct. Biol. 4, 10). The natively folded conformation (N) locates at the tip of the funnel.
Investigation of conformational energy landscape of proteins on a proteomic scale
With the advent of the postgenomic and proteomic era, we face new challenges and new opportunities in protein folding studies. Can we obtain information on the energetics and dynamics of proteins on a proteomic scale? Can we study the relationship between the energetics and function of proteins at a system level? To address these issues, we use proteolysis as a structural probe. Conventional biophysical approaches using spectroscopy and calorimetry allow us to study only one protein at a time. However, by using proteolysis and proteomics tools, we can study energetics of multitude of proteins in a proteome at the same time. Currently, we attempt to determine global stabilities and unfolding kinetics of proteins on a proteomic scale. This research will allow us to understand why some proteins are more stable than the others and how evolution has shaped the distribution of thermodynamic and kinetic stabilities of proteins in a proteome.
Identification of cellular drug targets by proteolysis
Energy landscapes of proteins are perturbed by interaction with drug molecules. By monitoring these changes on a proteomic scale using proteolysis, we can identify cellular targets interacting with drug molecules. The cellular targets of effective drugs are often unknown. Moreover, novel chemicals affecting cellular activities are discovered at a tremendous speed by chemical genetics. By knowing the cellular target, we can design better drugs based on the structure and screen other drug candidates in vitro. In addition, these drugs serve as research tools to control cellular functions of the target. The following diagram shows the concept of drug target identification using proteolysis.
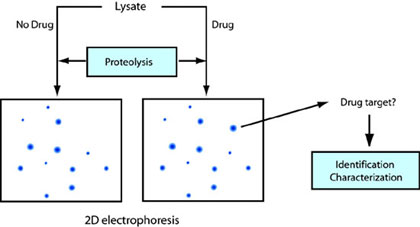
Investigation of kinetic and thermodynamic stability of membrane proteins and ligand binding to G-protein coupled receptors
In spite of the functional importance of membrane proteins, they have not been favorite research subjects in research on protein energetics. Membrane proteins are hard to express and purify. Therefore, conventional biophysical studies of membrane proteins are quite challenging. Using proteolysis as a structural probe, we are making a breakthrough in membrane protein research. Currently we aim to determine thermodynamic and kinetic stabilities of endogenous membrane proteins in cells without purifying them. This research will enhance our understanding of membrane protein energetics and dynamics significantly. Also, by measuring the change in stability by ligand binding, we are developing quantitative non-radioactive ligand binding assays for G-protein coupled receptors. Because G-protein coupled receptors are prominent drug targets, the novel assay methods will have a great impact in drug discovery and pharmacological research.
Chiwook Park
Visual Interpretation of the Meaning of kcat/KM in Enzyme Kinetics.
Journal of Chemical Education 99, 2556–2562 (2022).
Chen Chen and Chiwook Park
Chaperone Action of a Cofactor in Protein Folding.
Protein Science 29, 1667–1678 (2020).
Nathan W. Gardner, Sarah M. McGinness, Jainik Panchal, Elizabeth M. Topp, and Chiwook Park
A Cooperative Folding Unit as the Structural Link for Energetic Coupling within a Protein.
Biochemistry 56, 6555–6564 (2017).
Chen Chen, Jung-Hun Yun, Jae-Hoon Kim, and Chiwook Park
Effect of Circular Permutations on Transient Partial Unfolding in Proteins.
Protein Science 25, 1483–1491 (2016).
Mark W. Hinzman, Morgan E. Essex, and Chiwook Park
Salt Bridge as a Gatekeeper against Partial Unfolding.
Protein Science 25, 999-1009 (2016).
Nathan W. Gardner, Lyman K. Monore, Daisuke Kihara, and Chiwook Park
Energetic Coupling between Ligand Binding and Dimerization in E. coli Phosphoglycerate Mutase.
Biochemistry 55, 17110-1723 (2016).
Joseph R. Kasper and Chiwook Park
Ligand Binding to a High-Energy Partially Unfolded Protein.
Protein Science 24, 129-137 (2015).
Jonathan P. Schlebach, Nicholas B. Woodall, James U. Bowie, and Chiwook Park
Bacteriorhodopsin Folds through a Poorly Organized Transition State.
Journal of the American Chemical Society 136, 16574-16581 (2014).
Joseph R. Kasper, Pei-Fen Liu, and Chiwook Park
Structure of a Partially Unfolded Form of Escherichia coli Dihydrofolate Reductase Provides Insight into its Folding Pathway.
Protein Science 23, 1728-1737 (2014).
Youngil Chang, Jonathan P Schlebach, Ross A VerHeul, and Chiwook Park
Simplified Proteomics Approach to Discover Protein-Ligand Interactions.
Protein Science 21, 128-1287 (2012).
Pei-Fen Liu and Chiwook Park
Selective Stabilization of a Partially Unfolded Protein by a Metabolite.
Journal of Molecular Biology 422, 403-413 (2012).
Jonathan P Schlebach, Zheng Cao, James U Bowie, and Chiwook Park
Revisiting Folding Kinetics of Bacteriorhodopsin.
Protein Science 21, 97-106 (2012).
Pei-Fen Liu, Daisuke Kihara, and Chiwook Park
Energetics-based Discovery of Protein–Ligand Interactions on a Proteomic Scale.
Journal of Molecular Biology 408, 147-162 (2011).
Jonathan P Schlebach, Moon-Soo Kim, Nathan H Joh, James U Bowie, and Chiwook Park
Probing Membrane Protein Unfolding with Pulse Proteolysis.
Journal of Molecular Biology 406, 545-551 (2011). (Cover Article)
Youngil Chang and Chiwook Park
Mapping Transient Partial Unfolding by Protein Engineering and Native State Proteolysis.
Journal of Molecular Biology 393, 543-556 (2009).
Moon-Soo Kim, Jiao Song, and Chiwook Park
Determination of Protein Stability in Cell Lysates using Pulse Proteolysis and Western Blotting.
Protein Science 18, 1051-1059 (2009).
Yu-Ran Na and Chiwook Park
Investigating Protein Unfolding Kinetics by Pulse Proteolysis.
Protein Science 18, 268-276 (2009).
Chiwook Park, Sharleen Zhou, Jacquline Gilmore, and Susan Marqusee
Energetics-based Protein Profiling on a Proteomic Scale: Identification of Proteins Resistant to Proteolysis.
Journal of Molecular Biology 368, 1426-1437 (2007).
Chiwook Park and Susan Marqusee
Pulse Proteolysis: A Simple Method for Quantitative Determination of Protein Stability and Ligand Binding.
Nature Methods 2, 207-212 (2005).
Chiwook Park and Susan Marqusee
Probing the High Energy States in Proteins by Proteolysis.
Journal of Molecular Biology 343, 1467-1476 (2004).

