
Ph.D. - 1981 - University of California, San Diego
Postdoc - 1982-85 - Harvard University
Not Accepting Students
The research focus of the Post group is broadly described as investigations to understand the regulation and function of protein-protein interactions associated with cell signaling and viruses. Multi-dimensional, multinuclear NMR methods are used to determine 3-dimensional structure of protein complexes. Computational methods are used to study the mechanism of action of antiviral compounds, and the molecular mechanism for phosphotyrosine control of protein function in signal transduction.
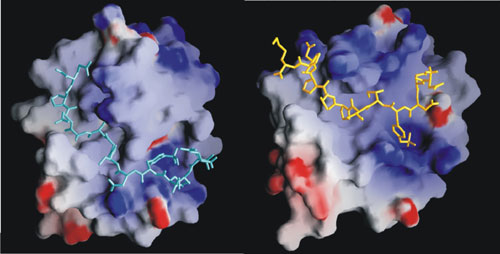
One research area of our group is the regulation of enzymatic activity and protein-protein association by tyrosine phosphorylation. Such regulation is central to cell signaling, and the transduction of chemical information from outside the cell to control intracellular processes that govern cell growth and differentiation. Protein tyrosine kinases are targets of high interest for cancer therapeutics and for drugs to treat immune diseases. We seek an atomic-level description to begin to understand the molecular basis of regulation. NMR structural studies and computational methods are being used to study how tyrosine phosphorylation controls enzymatic activity of Src-family kinases and other tyrosine kinases in B-cells. One aim is to unravel the molecular mechanism of activation. We are also actively engaged in developing new procedures to facilitate and improve the structure determination method by NMR.
Another area of research interest is the study of viral assembly processes using computational biology and high-resolution NMR methods. Results from molecular dynamics on human rhinovirus lead to a new proposal to explain the antiviral activity of WIN compounds against rhinovirus; binding of these compounds in an internal pocket of rhinovirus changes the viral capsid from a rigid-like protein shell to one that is more compressible and thus entropically stabilized. Similar approaches are now being applied to understand virus-receptor interactions and cell entry.
High resolution NMR spectroscopy is also used to elucidate the structure and dynamics of alphaviral and retroviral proteins. Specific interactions at a membrane interface between protein core particles and transmembrane glycoproteins necessary for proper budding of alphaviruses are being investigated. In addition, maturation of retroviruses requires specific protein-protein interactions of the gag polyprotein to form infectious particles. NMR structural studies were recently initiated on this system. Methods to obtain solution structures efficiently by a combined use of NMR data and computational tools are under development and should be useful in proteomic initiatives.
James R. (Jimmy) Ellegate (Graduate Student)
Vivek Govind Kumar (Post-Doctoral Research Associate)
Dhulika Ravinuthala (Graduate Student)
Rajarshi Roy (Post-Doctoral Research Associate)
Dan Xie (Graduate Student)
Z. Zhou, C. Patkar, J. Suk, M. Khaliq, L. Li, et al. 2008. Antiviral compounds discovered by virtual screening of small-molecule libraries against dengue virus E protein. accepted ACS Chem. Biol.
E. Ozkirimli, S. Singh, W.T. Miller, C.B. Post. 2008. An electrostatic network is functional in the long-range regulation of Src kinases. Protein Science (cover illustration), accelerated publication, Nov 2008.
Y. Zhang, H. Oh, R.A. Burton, J.W. Burgner, R.L. Geahlen, C.B. Post. 2008. Tyr130 phosphorylation triggers Syk release from antigen receptor by long-distance conformational uncoupling. Proceedings of the National Academy of Sciences 105:11760-11765.
V.M. Dadarlat, C.B. Post. 2008. Contribution of Charged Groups to the Enthalpic Stabilization of the Folded States of Globular Proteins. J. Phys. Chem. B 112:6159-6167.
R.A. Burton, S. Mattila, E.J. Taparowsky, C.B. Post. 2006. B-Myc: N-Terminal Recognition of Myc Binding Proteins. Biochemistry 45:9857-9865.
T.D. Groesch, F. Zhou, S. Mattila, R.L. Geahlen, C.B. Post. 2006. Structural Basis for the Requirement of Two Phosphotyrosine Residues in Signaling Mediated by Syk Tyrosine Kinase. Journal of Molecular Biology 356:1222-1236.
LI, Y., Zhou, Z. & Post, C. B. (2005) "Dissociation of an antiviral compound from the internal pocket of human rhinovirus 14 capsid," Proc. Natl. Acad. Sci., 102(21): 7529-7534.
L. Ma, C.T. Jones, T.D. Groesch, R.J. Kuhn and C.B. Post (2004) "Solution Structure of Dengue Virus Capsid Protein Reveals a New Fold," Proc. Natl. Acad. Sci., 101: 3414-3419.
C.B. Post (2003) "Exchange-transferred NOE spectroscopy and bound ligand structure determination," Curr. Opin. Struct. Biol., 13(5): 581-588.
V.M. Dadarlat and C.B. Post (2003) "Adhesive-cohesive model for protein compressibility, a new perspective on stability," Proc. Natl. Acad. Sci., 100(25): 14778-14783.

